7.2.2. Data management#
This section shows how to use these methods for data management:
We will use this sample kinematic acquisition of 39 bilateral markers with 1092 samples recorded at 50 Hz during tennis serve:
import kineticstoolkit.lab as ktk
import matplotlib.pyplot as plt
markers = ktk.read_c3d(ktk.doc.download("kinematics_tennis_serve.c3d"))[
"Points"
]
7.2.2.1. Adding data#
The TimeSeries loaded above has many markers, which makes lots of data to visualize and process:
markers.plot()
UserWarning [/Users/felix/Documents/git/kineticstoolkit_doc/src/kineticstoolkit/timeseries.py:4364] Only 40 of 156 lines have been plotted. Increase max_lines to plot more lines.
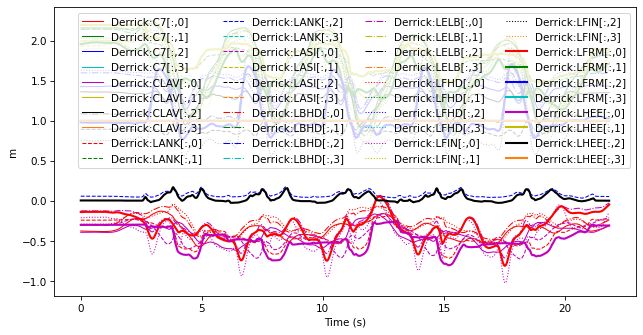
To keep only the data we need, it is often practical to make a copy of the TimeSeries with no data, and selectively add the data we need using ktk.TimeSeries.add_data:
# Copy the TimeSeries without its data
selected_markers = markers.copy(copy_data=False)
# Add the markers to this new TimeSeries
for key in ["Derrick:C7", "Derrick:T10", "Derrick:STRN", "Derrick:CLAV"]:
selected_markers = selected_markers.add_data(key, markers.data[key])
selected_markers.plot()
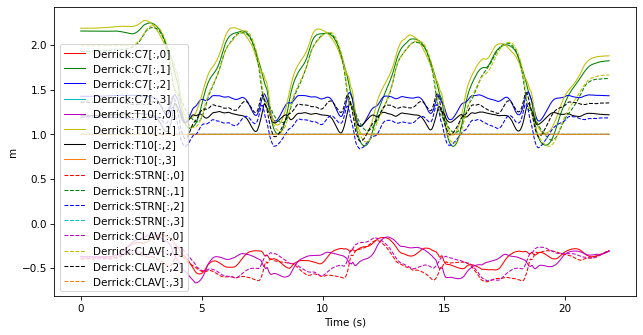
7.2.2.2. Removing data#
We could do the same by removing the superfluous data from the TimeSeries, using ktk.TimeSeries.remove_data:
# Copy the whole TimeSeries, with all data
selected_markers = markers.copy()
# Delete the unwanted markers from this new TimeSeries
for key in markers.data:
if key not in ["Derrick:C7", "Derrick:T10", "Derrick:STRN", "Derrick:CLAV"]:
selected_markers = selected_markers.remove_data(key)
selected_markers.plot()
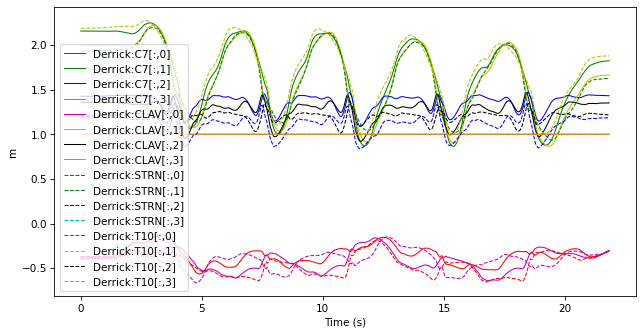
7.2.2.3. Subsetting#
As a shortcut method, we can also use ktk.TimeSeries.get_subset, which returns a TimeSeries with only a selection of data keys:
selected_markers = markers.get_subset(
[
"Derrick:C7",
"Derrick:T10",
"Derrick:STRN",
"Derrick:CLAV",
],
)
selected_markers.plot()

7.2.2.4. Renaming data#
Use the ktk.TimeSeries.rename_data to change the name of a data key:
selected_markers = selected_markers.rename_data("Derrick:C7", "C7")
selected_markers = selected_markers.rename_data("Derrick:T10", "T10")
selected_markers = selected_markers.rename_data("Derrick:STRN", "Sternum")
selected_markers = selected_markers.rename_data("Derrick:CLAV", "Clavicular")
selected_markers.plot()
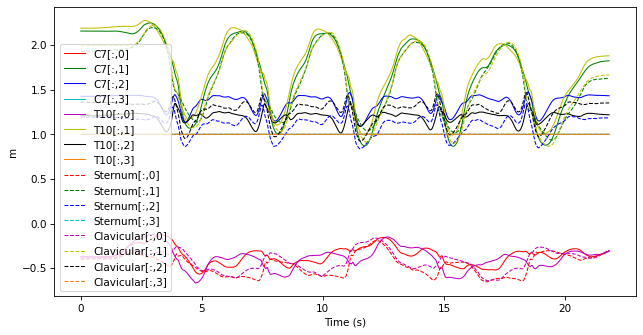
7.2.2.5. Merging data#
Let say we have two different TimeSeries, one for the left lower limb and one for the right one:
left_lower_limb = markers.get_subset(
[
"Derrick:LANK",
"Derrick:LHEE",
"Derrick:LKNE",
"Derrick:LTOE",
]
)
right_lower_limb = markers.get_subset(
[
"Derrick:RANK",
"Derrick:RHEE",
"Derrick:RKNE",
"Derrick:RTOE",
]
)
plt.subplot(1,2,1)
left_lower_limb.plot(legend=False)
plt.subplot(1,2,2)
right_lower_limb.plot(legend=False)

To merge these TimeSeries, we could simply use the ktk.TimeSeries.add_data method as we did above:
merged = left_lower_limb.copy()
for key in right_lower_limb.data:
merged = merged.add_data(key, right_lower_limb.data[key])
merged.plot(legend=False)
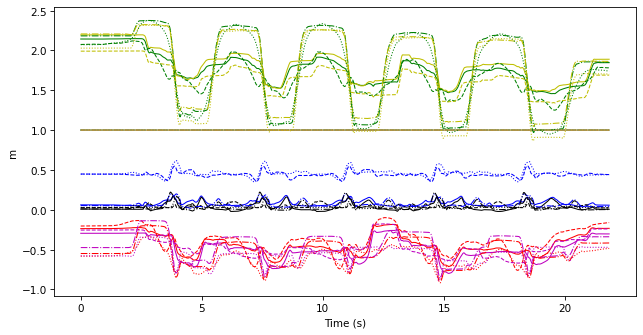
As a shortcut method, we can also use ktk.TimeSeries.merge:
merged = left_lower_limb.merge(right_lower_limb)
merged.plot(legend=False)

Using ktk.TimeSeries.merge has several other benefits, such as merging events, information, and even resample TimeSeries that may have been recorded at different sampling rates.
